- What version of pylinac are you using (
pylinac.__version__)?
pylinac 3.24.1
- I’ve read the documentation for this module? (Yes/No/Doesn’t Exist)
Yes thoroughly. I notably tried applying a filter
Hi everyone,
I have a little issue, and I would be happy to learn if anyone has a solution ![]()
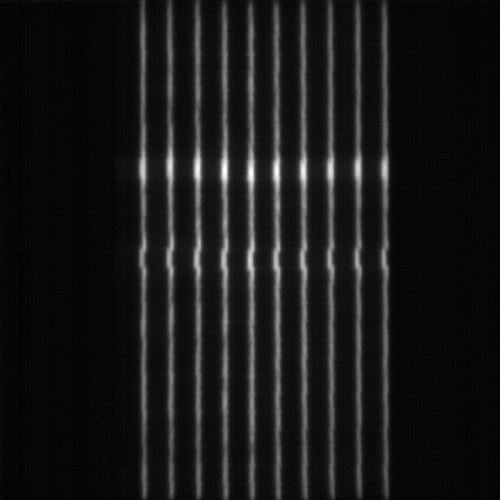
I have a picket Fence DICOM file that looks like this :
I can analyse it just fine using this code snippet:
from pylinac import PicketFence
from pylinac.picketfence import MLC
path =r "mypath.dcm"
pf = PicketFence(filename=path, mlc=MLC.MILLENNIUM) # Millennium is the default
pf.analyze()
pf.plot_analyzed_image()
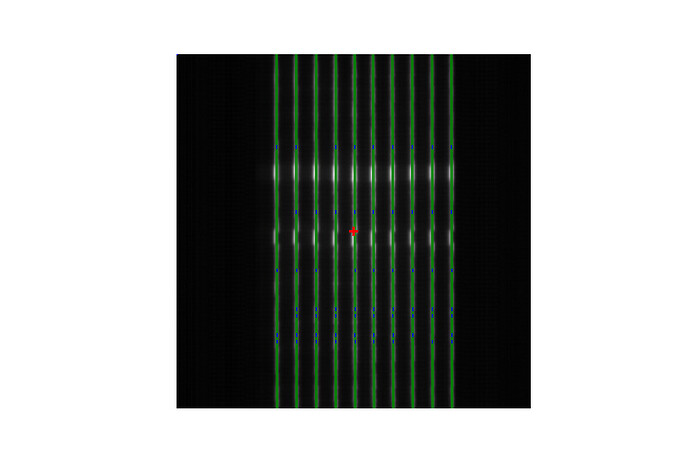
This returns:
However this Picket Fence image is from a Halcyon using both proximal and distal leaves, so I tried with this code:
from pylinac import PicketFence
from pylinac.picketfence import MLCArrangement
mlc_setup=MLCArrangement([(57,5)])
#There are 28 leafs for the distal and 29 for the proximal
#Each leaf has a width of 10 mm with an offset of 5 mm between the distal and proximal leaves
path=r"mypath.dcm"
pf = PicketFence(filename=path, mlc=mlc_setup)
pf.analyze()
pf.plot_analyzed_image(leaf_error_subplot=False) #I get an error if I put it on True
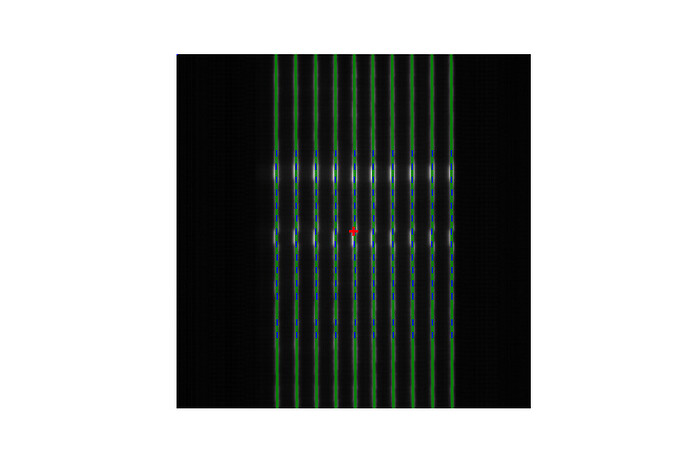
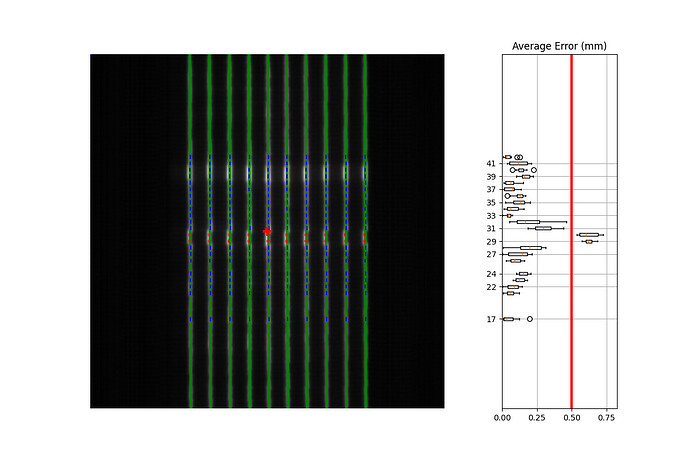
But I get this result:
If I use pf = PicketFence(filename=path, mlc=MLC.HALCYON_PROXIMAL)
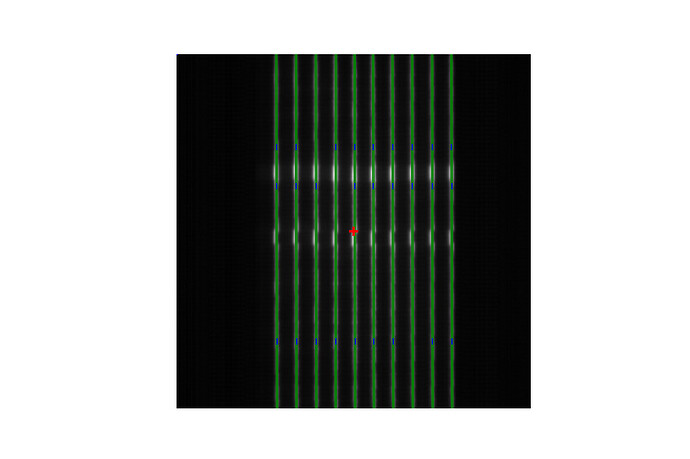
it doesn’t detect the error:
And it doesn’t work either with
pf = PicketFence(filename=path, mlc=MLC.HALCYON_DISTAL)
So what do you think ? Should I create 2 Picket Fence images, one with the distal leaves and one with the proximal ? I’m not sure that it’s the best practice because this isn’t what we use on patients. What do you do in your clinic ? I know this is a commonly asked question (it was also asked in the latest French Radmachine webinar) but I’m unsure how to proceed.
Regards to whoever stuck to the end of this long post,
Safir